KIRHub is an interactive web-based platform designed for researchers and clinicians
to explore the relationship between clinically-approved kinase inhibitors and their effects on
both wild-type and oncogenic kinase variants. The application is fully browser-based,
requiring no data uploads, ensuring ease of use and accessibility.
Please cite the publication below for use of this tool.
Kinase Inhibitor Database Overview
KIRHub: User Guide
KIRHub is a browser-based platform for exploring activity profiling data of clinically approved kinase inhibitors. It provides three interactive modules: (i) identifying drug(s) to inhibit wild-type kinases, (ii) identifying drug(s) to inhibit mutant kinases, and (iii) exploring kinase essentiality across cancer lineages. Each module supports filtering, sorting, visualization, and CSV/PNG downloads—designed to surface polypharmacology and repurposing hypotheses efficiently.
Quick Start Recommended
- Choose a module: Use the three large buttons on the top of the app to open either Wild-Type Kinases, Mutant Kinases, or Cancer Lineages.
- Search/select: Start typing a kinase/mutation; the list filters dynamically. Exact matches are best (e.g.,
ABL1;BRAF(V600D);FGFR1OP/FGFR1). - Review ranked drugs: The table shows the top candidates by % inhibition (and selectivity via Gini Scores). Use column headers to sort; use the download button to export results.
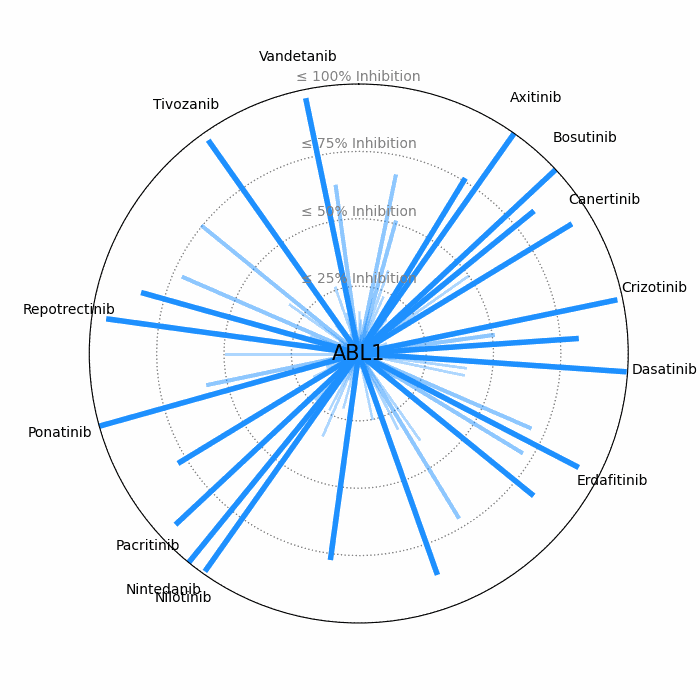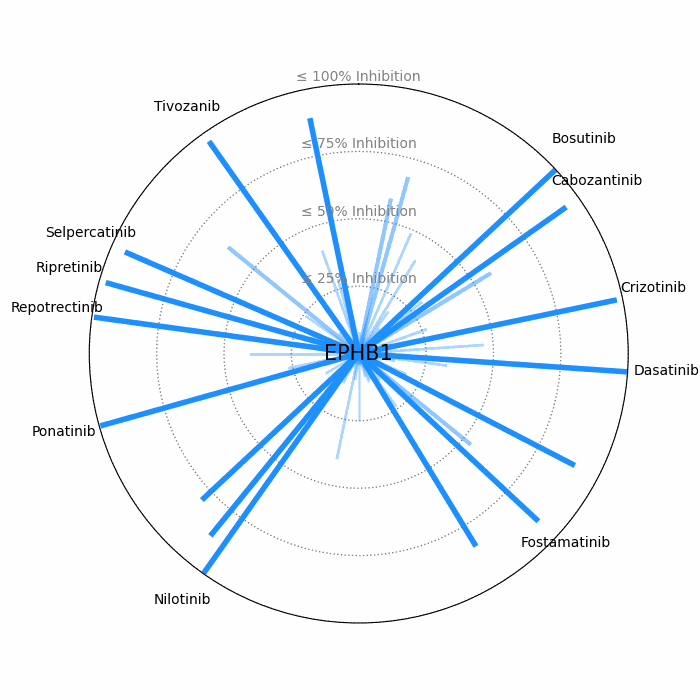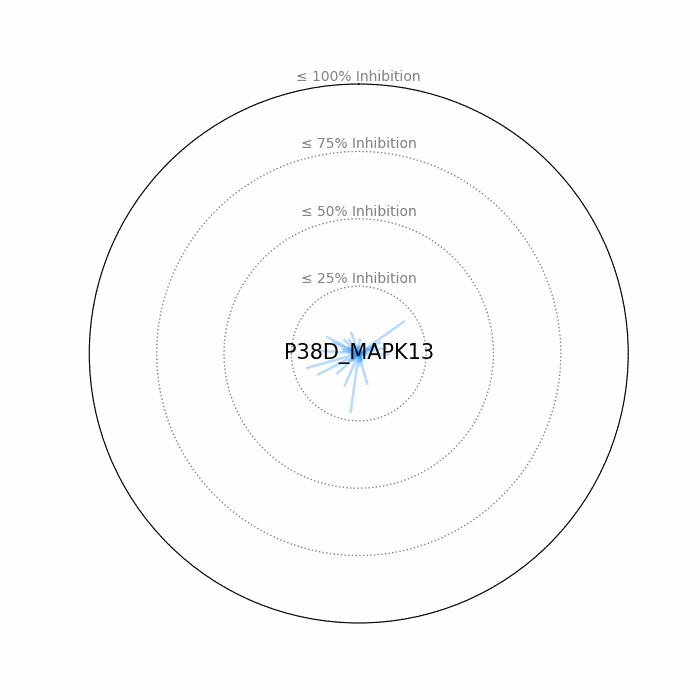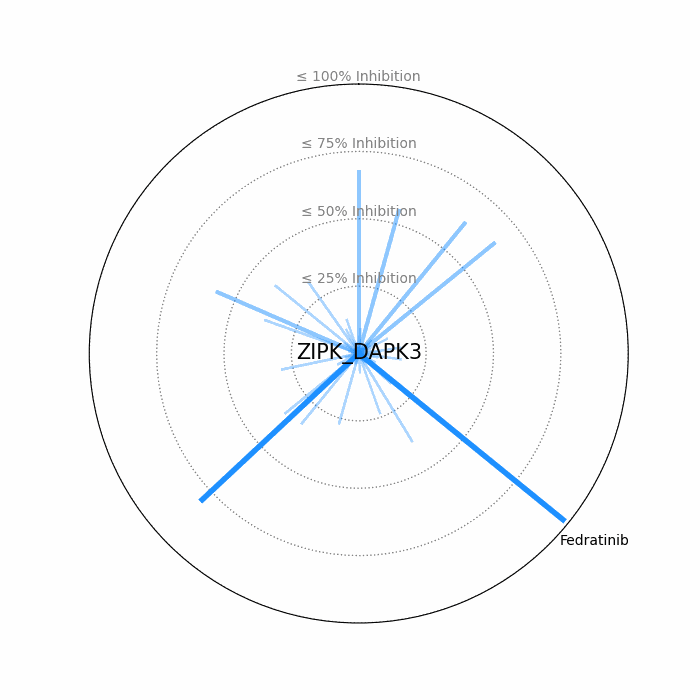
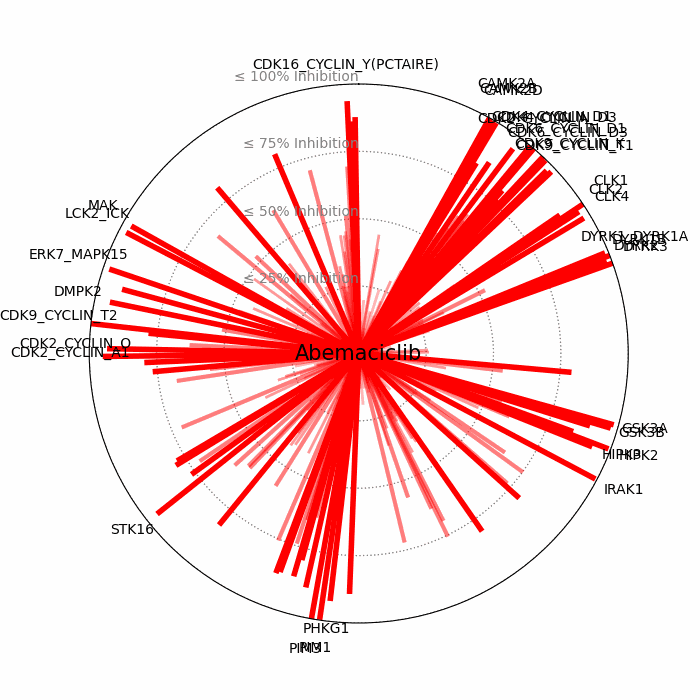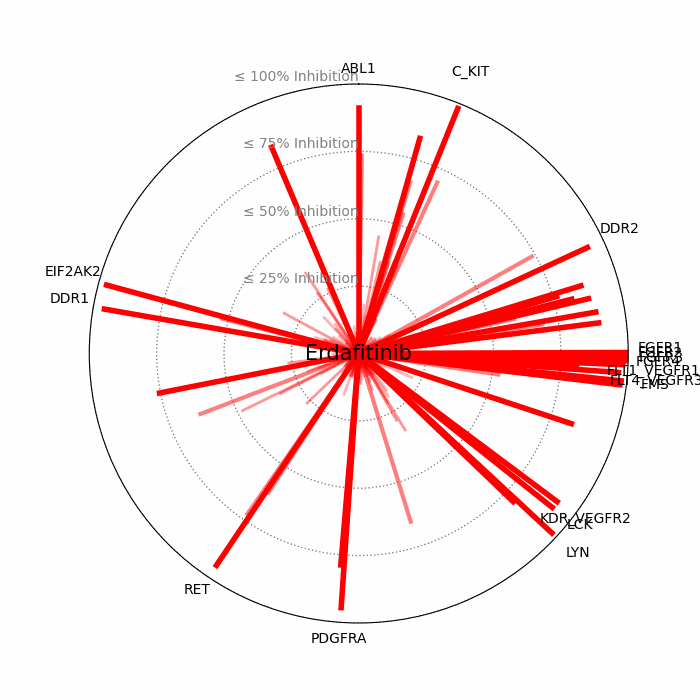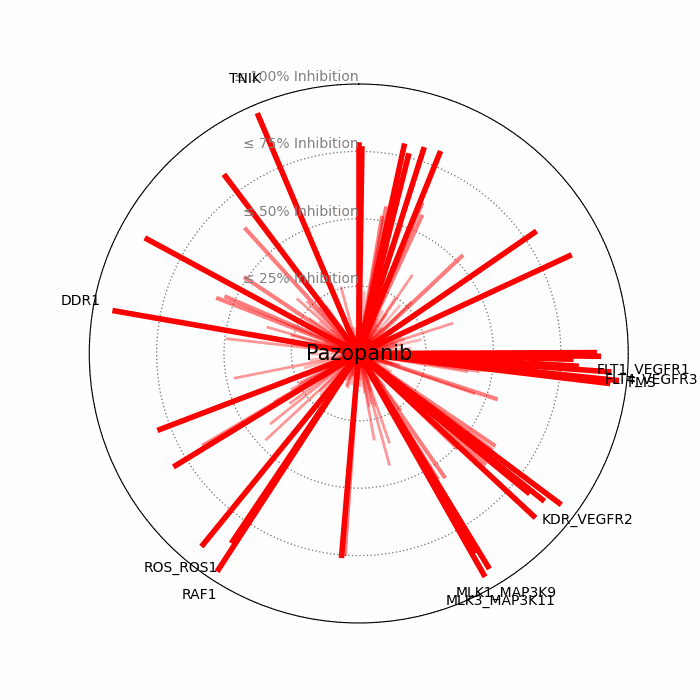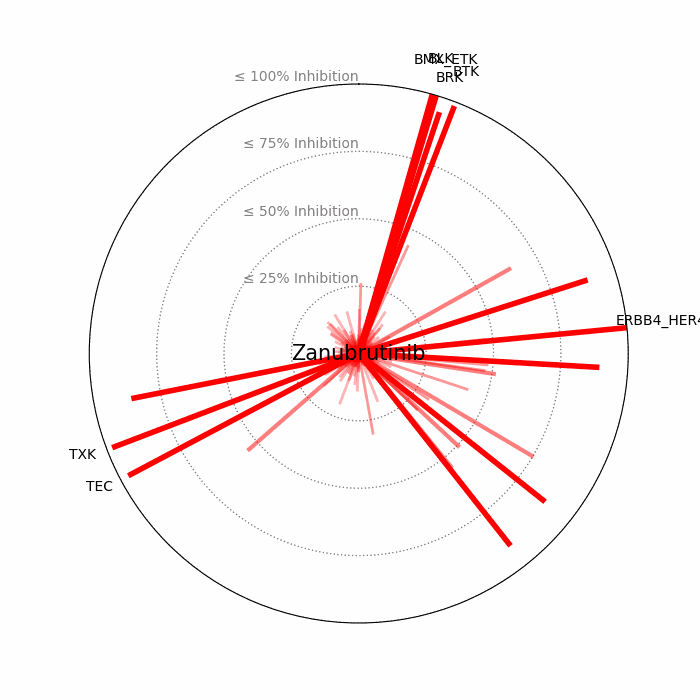
- Open visuals: Radar plots display per-drug inhibition distribution; KISS vs Inhibition scatters highlight drugs balancing on-target inhibition and off-target burden.
- Dual targets (optional): Add a second kinase/mutation to see drugs that score highly for both (intersection of top candidates).
Module 1 — Identifying Drug(s) to Inhibit Wild-Type Kinases
Goal: Prioritize inhibitors for a wild-type kinase (or dual kinase or paralog pair).
- Select a kinase (e.g., P38A/MAPK14, ABL2/ARG, AURKA).
- Optionally select a second kinase/paralog to co-prioritize.
- Review ranked drugs by % inhibition and Gini. Download table as CSV; download Radar/KISS figures as PNG.
- KISS vs Inhibition: Points in the upper-right (higher KISS and higher % inhibition) are favorable.
Module 2 — Identifying Drug(s) to Inhibit Mutant Kinase(s)
Goal: Find clinically approved inhibitors with high inhibition for a selected mutant kinase (optionally a combination).
- Select a mutant (e.g.,
BRAF(V599E),ALK/TFG Fusion,FGFR1(V561M)). - Optionally select a second mutant to prioritize drugs effective on both.
- Interpret the table: Inhibition (%) is computed as
100 − residual activityat 1 μM. Gini approximates selectivity (higher is more selective). - Use the download button to export ranked results (CSV).
- Open Radar Plot(s) for a quick visual of drug inhibition patterns across candidates.
Module 3 — Exploring Kinase Essentiality Across Cancer Lineages
Goal: See where a kinase is essential across primary and nested lineages (DepMap-derived). “Essential” is a kinase that has a gene effect score of less than -0.5 in a particular cancer lineage and its sub-lineages. These gene effect scores data are obtained from the DepMap Portal website.
- Select a kinase; the table lists lineage counts and percentages where it’s essential.
- Use the bar chart to compare total essential counts by Lineage 1.
- Export the table or bar chart (CSV/PNG).
Assay Metadata (Reaction Biology) — Downloadable Files
These files describe assay setup for each kinase used in biochemical profiling: whether the full-length protein or a domain fragment (with residue range when available) was assayed; the expression system (e.g., baculovirus/Sf21) and affinity tag (e.g., His/GST); and the general substrate (e.g., Abltide, Crosstide, Casein+Mn, etc.). This context addresses concerns about regulatory domains affecting activity/folding and substrate choice shaping readouts.
How to Read the Visuals
- Inhibition (%): Higher is better and equals
100 − residual activityat 1 μM. - Radar Plot: Each spoke is a candidate drug; longer spokes = greater inhibition. The concentric inhibition guides (≤25/50/75/100%) help judge magnitude quickly.
- KISS Score: Balances on-target effect (geometric-mean inhibition on the chosen kinase set) against off-target burden. Use alongside % inhibition to find selective, potent options.
- Gini Score: Higher score suggests more selective inhibition profile across kinases.
Submit a New Dataset to KIRHub
If you have a kinase inhibition dataset or compound screen that you would like to see included in KIRHub, please follow the steps below.
Step 1 — Validate Dataset Format
Upload a .csv or .xlsx file to check whether it matches the required KIRHub format. No data are stored or analyzed automatically.
Step 2 — Contact the Authors to Submit a New Dataset
Once your file passes validation, please submit it by contacting the authors directly either via email or using the citation link below. Include a short description of the dataset and its source.
Email the KIRHub authorsCitation:
Required File Format
- Column 1: Compound (aliases in parentheses allowed; e.g. Lapatinib (GW572016))
- Column 2: CAS (or other unique compound identifier; e.g. 231277-92-2)
- Column 3: Dose (e.g. 0.5, 1uM; no spaces) in uM only
- Columns 4+: HGNC kinase names (all caps, no whitespace; e.g. AURKA, MAPK1, EGFR). Same for mutants; eg. FGFR3(V555M), EGFR(D746_750_C797S)
- Values: Numeric residual activity values (0–100) or NA only, where 0 indicates complete inhibition and 100 indicates no inhibition relative to control.
- Characters: Standard US keyboard characters only (e.g. use u instead of Greek μ)
Tips, Caveats, and Troubleshooting
- Search strategy: Prefer exact names first (e.g.,
P38A/MAPK14) before abbreviations. For variants, include mutation (e.g.,BRAF(V599E)) or '/' fusion tag. - Dual-target logic: By default, the app intersects top-ranked lists per target; if the intersection seems too strict, try each target separately to see near-misses.
- Units: Inhibition values are computed at 1 μM experimental dose.
- Downloads: Look for the Download Table and Download Plot buttons below each output.
- Empty/NA results: If no values appear, try a different spelling or a broader kinase family member (paralog).
Please cite the following publication for use of this tool:
Learn more
To learn more about the kinase inhibitor profiling efforts in the Gujral Lab, including the network pharmacology approaches underlying this resource, please click here
To learn how these insights are being applied to rare cancers through functional precision oncology, visit TRACER.
Note: Kinase inhibition data are the property of Reaction Biology Corporation. Proper acknowledgment is required for download/use/publication. For questions: info@reactionbiology.com.
